Researchers use visualization tools to render 3-D medical images
Researchers use visualization tools to render 3-D medical images
By Andrew Wilson, Editor at Large
In the past, physicians depended as much on physical evidence as laboratory data to diagnose patients. Now, however, medical imaging systems have evolved to the point where it is possible to see certain anatomical structures with very fine detail. High-resolution computed-tomography (CT) systems, for example, now allow organs to be visualized at resolutions approaching the submillimeter level. Better still, fusion of data from different imaging modalities can yield images of the structural and functional relationships of such organs.
As such systems become more sophisticated, the data they produce increase dramatically. To understand these data, physicians and researchers need to visualize images of theanatomy in a lifelike way. To do so, they are turning to three-dimensional (3-D) visualization software to examine, explore, and precisely pinpoint problem areas before determining final treatment strategies.
Choosing a package
The challenge is to create easy-to-use graphical-rendering and analysis tools. To produce fast, powerful visualization technology, medical application developers are turning to software-development tools that convert data from a variety of medical instruments into interactive visual environments.
Providing such capability does, however, come at a price. Because the computational requirements and data sets are very large, most third-party packages, such as IDL from Research Systems (Boulder, CO), IMOD from the University of Colorado (Boulder, CO), and the Visualization Data Explorer from IBM (White Plains, NY), require graphics workstations or UNIX-based hosts.
In choosing a visualization package to perform a specific imaging function, developers must carefully evaluate a number of factors. These include hardware requirements, data types and file formats supported, visualization and viewing techniques used, image-analysis capability, data fusion, and virtual reality. Developers choosing the Voxel View from Vital Images (Minneapolis, MN), for example, must run the 3-D volume-rendering software on either the R5000SC or R10000SC O2 graphics workstation from Silicon Graphics (Mountain View, CA). At a price of $35,000 including the graphics workstation, VoxelView allows developers to visualize, analyze, and manipulate 3-D image data, highlight regions of interest, control transparency, color, and lighting, and create digital animations.
Like VoxelView, SciAn, developed at the Supercomputer Computations Research Institute at Florida State University (Tallahassee, FL), is a UNIX-based scientific visualization and animation package that runs on both Silicon Graphics and IBM RS/6000 workstations. Version 1.1 of the software, currently in beta test, supports a number of different data types. These include scalar and vector fields, structured and nonstructured grids, and datasets that vary over time. In operation, the software can show any number of visualizations on screen at any one time. Viewpoints of different visualizations can be linked, enabling comparison of data from observation and simulation.
Perhaps the most important consideration in choosing a visualization package is the type of file format supported. Because data formats from MRI, PET, and CT equipment differ, software packages such as MEDx from Sensor Systems (Sterling, VA) support a number of different file formats including data types from Picker MRI, Siemens PET, and GE Signa MRI scanners. Also running on a variety of UNIX workstations including the 300 Series from HAL Computer Systems (Campbell, CA), MEDx software is designed to display both 2-D and 3-D data under X Windows. Release 2.0 of the software includes a functional image-analysis module to normalize, detect, and correct for head motion and an automated 3-D registration algorithm developed by Roger Woods at UCLA (Los Angeles, CA) that can register noncoplanar data sets.
Image analysis
Once loaded into memory, a number of image-analysis techniques can enhance or measure image data. Typical image-processing routines include 3-D matrix operations, linear and adaptive histogram functions, 2-D and 3-D fast-Fourier-transform (FFT) and deconvolution routines, and fusion of multimodal images. These are used to both enhance and extract data from the images. To make quantitative measurements on image data, many off-the-shelf packages also have image-analysis functions. These include plotting of line and trace profiles, selection and automatic plotting of regions of interest, and surface and volume area measurement.
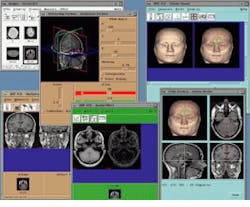
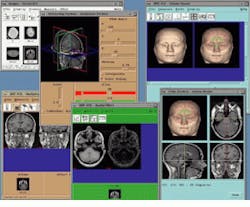
As part of its Analyze AVW package, the Biomedical Imaging Resource at the Mayo Clinic (Rochester, MN) includes a number of techniques to analyze, measure, and display 3-D image data. Running under UNIX, Analyze allows multiple volume images to be simultaneously accessed and processed by multiple programs in a multi-windows interface (see Fig. 1). AVW functions are available to developers in two forms: either as a C-callable library of functions or as extensions to a Tcl/Tk software toolkit. The C-callable library permits the AVW functions to be compiled and linked to software systems both statistically and dynamically. The Analyze package is currently available from CNSoftware (Stewartville, MN).
Visualization techniques
To display 2-D and 3-D images, visualization techniques allow data to be viewed as orthogonal or oblique sections, as volume or surface- rendered objects, or as a movie of volume or surface-rendered images. While orthogonal sections can provide a general overview of image data sets and allow the operator to visually compare images, oblique sections can extract slices at any arbitrary orientation within a volumetric data set. Better still, new data sets with slices parallel to a selected oblique angle can be generated after resampling and interpolation.
As part of a drug-design program at Bristol-Meyers Squibb Pharmaceutical Research Institute (Princeton, NJ), Dr. Ron Behling is using MRI data sections to study problems in drug discovery and design. To do so, Behling is using PV-Wave from Visual Numerics (Houston, TX) running on a Silicon Graphics workstation. "Although there are no standard data formats in MRI," says Behling, "the command language of PV-Wave allows virtually any data types to be read." Using various PV-Wave functions, such as FFTs and curve fits, Behling is developing custom procedures to study diffusion rates in tissues.
In many cases, it is useful to observe the 2-D structure of organs in the body. To do so, many visualization packages incorporate volume rendering, a technique used for 3-D visualization of structures such as the heart and lungs. Produced by projecting rays from multiple angles of view through the volumetric data set, rendered objects can then be scaled and rotated about x, y, and z axes to produce animated 3-D displays.
"Tumors are often very irregular and positioned close to other structures. Three-dimensional rendering is therefore very important when you need to visualize the relative positions of these structures," says Zachary Leber, a ra diotherapy-development manager with Radionics Software Applications (Burlington, MA). Because of this, Radionics developed Xknife using the Application Visualization System (AVS) from Advanced Visual Systems (Waltham, MA). In operation, Xknife converts data from CT and MRI scans that show the location of a lesion into 3-D images that show its shape and position relative to optic nerves and other important structures (see Fig. 2). Physicians use Xknife to plan noninvasive treatment of intercranial abnormalities, sparing patients the expense, pain, and risk associated with open-brain surgery.
Surfaces and motion
In many medical applications, however, it is not necessary to understand data within 2-D data sets or to visualize data in three dimensions. In the manufacture of prostheses, for example, physicians only need to understand how the surfaces of structures are composed. Consequently, most software packages incorporate algorithms to allow structures such as bones to be displayed as 3-D shaded surfaces. To create 3-D shaded surface displays of structures, surface-rendering techniques often allow setting multiple threshold ranges to identify multiple objects within a volume. In this way, when two or more objects are to be displayed together, each can be assigned unique colors.
"The construction and depiction of surfaces from volume data are important in the manufacture of titanium prostheses for cranioplasty," says Dr. Jonathan Carr at the University of Cambridge (Cambridge, England). Previously with Christchurch Hospital (Christchurch, New Zealand), Carr developed a method for designing cranial implants to repair defects in the skull.
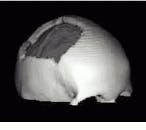
Skull surfaces were extracted from CT data by ray-tracing iso-value surfaces. By using a tensor product b-spline interpolation in the ray-caster, ripples in the surface data due to large spacing between CT slices were reduced. Associated depth maps characterize the defect regions that need to be repaired. Using radial basis functions, the surface of the skull was then interpolated and the fitted surface used to produce CNS milling instructions to machine a mold in the shape of the surface from a block of hard plastic resin. A cranial implant is then formed by pressing flat titanium plate into the mold using a hydraulic press (see Fig. 3).
In many imaging applications, physicians need to see the geometric relationships between anatomical structures. To do so, software packages may include techniques that allow multiple surface or volume rendered images to be rapidly displayed, thus producing the illusion of motion. This is especially useful in studying the heart, for example, where sequences of moving images can indicate loss of muscle function.
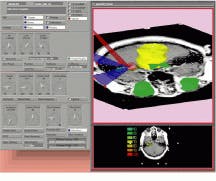
VIDA from the department of radiology at the University of Iowa College of Medicine (Iowa City, IO) allows developers to display images in several ways. Written in C and running under UNIX, VIDA has been developed for the manipulation, display, and analysis of multidimensional data sets. Available programs in VIDA include orthogonal and oblique sectioning, volume and surface rendering, ROI analysis, and interactive image segmentation (see Fig. 4). Once composed, a module called MovieViewer allows animation techniques to create the illusion of motion. And, because the software is modular, developers can rapidly integrate new programs.
Fusion and virtual reality
MRI images can show the extent of a tumor in relation to other soft tissues but not necessarily whether the tumor has invaded bone structures. CT images, on the other hand, can show bone structures but may give a relatively poor image of the soft tissue. Side by side images from the same patient can be difficult to interpret as the exact correspondence of the images is not known. Because of this, researchers are looking to image-fusion techniques to combine both data sets in single, composite images, removing the problems associated with side-by-side evaluation of images from different medical imaging modalities.
At Foster Findlay Associates (FFA; Newcastle upon Tyne, England), researchers are developing CT/MRI image-fusion techniques using CImages 3-D. Running on UNIX and PC-based platforms, Cimages3-D is the result of a three-year development program between FFA and Shell Research (The Netherlands). As a development toolkit, the package`s image-processing functions include image enhancement, segmentation, texture analysis, and ROIs within images. The package supports multiple image formats including CT, MRI, and PET and is supplied with an API that allows developers to build applications by combining image sequences.
Simulation and planning of craniofacial surgery is also the aim of the Virtual Environment for Reconstructive Surgery (VERS) project. A collaboration between the Biocomputation Center at NASA Ames Research Center (Moffett Field, CA) and the department of reconstructive surgery at Stanford University (Stanford, CA), the project aims to create a virtual environment where a surgeon can plan and simulate an operation (see Fig. 5). Using the Immersive Workbench from FakeSpace (Mountain View, CA), the project was demonstrated at last year`s SIGGRAPH by Silicon Graphics and Fakespace.
Computational power
Designers looking to incorporate 3-D visualization packages into their next generation of imaging equipment now have many off-the-shelf image-processing packages to choose from. In specifying such packages, developers should be careful to choose modular packages based on open standards that run on a number of hardware platforms.
Although most currently available packages do not run under Windows NT, it is highly likely that this will be the operating system of choice for many systems in the future. Better yet, because most packages available today are written in C and run under UNIX, it will be a simple task for software developers to create Internet-based, Java-enabled software that will run under multitasking, multiuser operating systems. When this is accomplished, sophisticated image-processing routines such as 3-D volumetric display, data fusion, and virtual-reality techniques will become as common as an aspirin.
FIGURE 1. Analyze AVW software from the Biomedical Imaging resource at the Mayo Foundation (Rochester, MN) permits simultaneuos use of multiple imaging tools assigned to any particular volume image or groups of volume images and other data elements.
FIGURE 2. Xknife software from Radionics Software Applications was developed using the Application Visualization System from Advanced Visual Systems. The Xknife user interface was built using AVS widgets and viewers and is shown with an MRI scan texture-mapped in the 3-D display.
FIGURE 3. To develop titanium cranial implants, ray-tracing is used to depict bone structures within a stack of CT data slices (A). Radial basis functions then fit a surface to the depth-map corresponding to the rendered view of the defect (B). A computer milling machine then produces a mold of the shape of the defect (C) that is used to construct a flat titanium plate that is screwed into the skull (D and E).
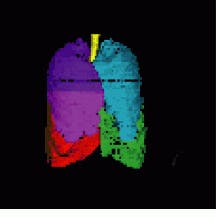
FIGURE 4. After gathering image data on an Imatron C-150 electron-beam CT scanner, color-coded images of the lungs were visualized by VIDA software from the division of physiologic imaging at the department of radiology at University of Iowa College of Medicine. The lung was segmented using VIDA`s segmentation module and displayed using the VIDA surface-display module.
FIGURE 5. Using the Immersive Workbench from Fakespace, physicians can interactively view and operate on three-dimensional images of patient data.