Convolutional neural network detects COVID-19 from chest radiography images
Linda Wang
Those keeping up with the news know that coronavirus testing helps track and stop the spread of the virus. In countries like the United States, testing remains behind where it should be, largely as a result of lack of tests and the tremendous burden placed on the national healthcare system. A new, open-source deep learning-based COVID-19 testing method offers hope in the form of an accurate, fast technique.
Caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the COVID-19 pandemic continues to devastate the health and well-being of the global population. While the main screening method— polymerase chain reaction (PCR) testing—detects SARSCoV-2 RNA from respiratory specimens reliably, it is a time consuming, laborious, and complicated manual process in short supply. Chest radiography imaging like X-ray or computed tomography (CT) provide radiologists visual indicators associated with the SARS-CoV-2 viral infection, as early studies show patients present abnormalities in chest radiography images characteristic of those infected with COVID-19, with some suggesting that radiography examination could be used as a primary tool for COVID-19 screening in epidemic areas.
While radiography exams can be done quicker and are more available given the prevalence of chest radiology imaging systems in modern healthcare systems, the need for expert radiologists to interpret the radiography images creates a bottleneck due to the subtle visual indicators present in the images.
While numerous artificial intelligence (AI) systems based on deep learning offer promising results in terms of accurately detecting COVID-19 via radiography imaging most or all of these are closed source and unavailable to the research community and for public use to build upon for deeper understanding and extension of these systems.
To that end, researchers from the University of Waterloo and the AI firm DarwinAI developed COVID-Net, a deep convolutional neural network tailored for the detection of COVID-19 cases from chest radiography images. The initial network design prototype aims to make one of the following four predictions: a) no infection (normal), b) bacterial infection, c) non-COVID viral infection, and d) COVID-19 viral infection. These possible predictions were chosen because results can aid clinicians to better decide not only who should be prioritized for PCR testing for COVID-19 case confirmation, but also which treatment strategy to employ, depending on the cause of infection, since each type of infection requires a different treatment plan. Figure 1 shows example chest radiography images of each of the four categories of infection.
Network architecture and dataset generation
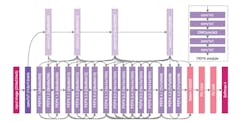
Figure 2 shows the proposed COVID-Net network architecture (available publicly for open access at bit.ly/VSD-COVID). The architecture makes heavy use of a lightweight residual projection-expansion-projection-extension (PEPX) design pattern, which consists of:
· First-stage Projection: 1 x 1 convolutions for projecting input features to a lower dimension,
· Expansion: 1 x 1 convolutions for expanding features to a higher dimension different than that of the input features,
· Depth-wise Representation: efficient 3 x 3 depth-wise convolutions for learning spatial characteristics to minimize computational complexity while preserving representational capacity.
· Second-stage Projection: 1 x 1 convolutions for projecting features back to a lower dimension, and
· Extension: 1 x 1 convolutions that finally extend channel dimensionality to a higher dimension to produce the final features.
Used to train and evaluate COVID-Net, the COVIDx dataset comprises 16,756 total chest radiography images across 13,645 patient cases. To generate the dataset, the team combined and modified two different publicly available datasets: COVID chest X-ray dataset and Kaggle chest X-ray images (pneumonia) dataset. These datasets were chosen because both are open source and accessible to the general public and research community, and as these datasets grow, so too will COVIDx.
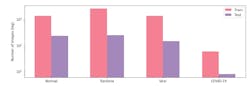
Figures 3 and 4 show the distribution of images and patient cases amongst different infection types. The most noticeable trend is the limited amount of COVID-19 infection cases and associated radiography images, which reflects the scarcity of COVID-10 case data available in the public domain, but also highlights the need to obtain more COVID-19 data as more case data becomes available to improve the dataset.
More specifically, the COVIDx dataset contains only 76 radiography images from 53 COVID-19 patient cases. For chest radiography images with no pneumonia, and non-COVID19 viral pneumonia, significantly more patient cases and corresponding radiography images exist. Images also includes a total of 8,066 patient cases who have no pneumonia (i.e., normal), and 5,526 patient cases with non-COVID19 pneumonia. Dataset generation scripts for constructing the COVIDx dataset are available at bit.ly/VSD-COVID.
Pretraining for COVID-Net was done using the ImageNet dataset, then trained on the COVIDx dataset using the Adam optimizer with a learning rate policy where the learning rate decreases when learning stagnates for a period. The following hyperparameters were used for training: learning rate=2e-5, number of epochs=10, batch size=8, factor=0.7, patience=5. The proposed COVID-Net was built and evaluated using the Keras deep learning library with a TensorFlow backend.
Implementation details and results
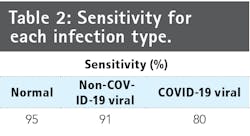
Quantitative and qualitative analysis methods evaluate the detection performance and decision-making behavior of COVID-Net. For quantitative analysis, the team computes test accuracy, sensitivity, and positive predictive value (PPV) for each infection type on the COVIDx dataset. Table 1 shows test accuracy, architectural complexity (number of parameters), and computational complexity (number of multiply-accumulation [MAC] operations). COVID-Net strikes a good balance between accuracy and computational complexity by achieving 92.4% accuracy while requiring 2.26 billion MAC operators for case prediction.
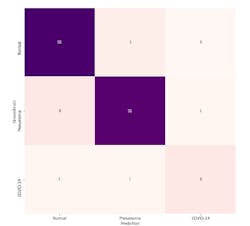
Studying the sensitivity and PPV for each in infection type shows the current COVID-Net limitations (Tables 2 and 3, along with a confusion matrix in Figure 5). COVID-Net shows 80% sensitivity for COVID-19 cases, an important factor since the software aims to minimize the number of missed COViD-10 patient cases. While quite promising, the number of COVID-19 patient cases available is limited compared to other infection types in COVIDx, and as such, effectiveness will improve as more COVID-19 patient cases become available. Additionally, COVID-Net achieves 88.9% PPV, indicating several false positive detections. Figure 5 shows two patients with bacterial infections that were misidentified as having COVID-19 viral infections. While this increases the burden of the healthcare system due to the need of additional PCR testing, it may be a reasonable trade off given the few false negative COVID-19 detections.
Furthermore, results show far fewer PPV for non-COVID-19 viral infection cases than for other infection types. Sensitivity for no infection cases is noticeably lower than other infection types. Results show that COVID-Net performs well in detecting COVID-19 cases from chest radiography images. Still, collecting additional data as well as improving the underlying training methodology to generalize better across such scenarios offer room for improvement.
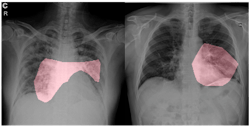
For qualitative analysis, the team explored how COVID-Net makes predictions by leveraging GSInquire—a proprietary technique from DarwinAI that provides insights into how deep neural networks reach decisions. Figure 6 shows critical factors identified in sample COVID-19 chest radiography images. COVID-Net identifies localized areas within the lungs and the chest radiography images as being critical factors in determining whether a radiography image shows a patient with a SARS-CoV-2 viral infection, as shown in red in Figure 6. The ability to interpret and obtain insights into how the proposed COVID-Net detects COVID-19 infections is important for three reasons:
· Transparency. By understanding the critical factors leveraged in COVID-19 case detection, the predictions made by the proposed COVID-Net become more transparent and trustworthy for clinicians to leverage during their screening process to aid them in making faster yet accurate diagnosis.
· New insight discovery. The critical factors leveraged by the proposed COVID-Net could potentially help clinicians discover new insights into the key diagnostic visual indicators associated with SARS-CoV-2 viral infection, which they can then leverage to improve screening accuracy.
· Performance validation. By understanding the critical factors being leveraged in COVID-19 case detection, one can validate that the proposed COVID-Net is not relying on improper information to make decisions (e.g., erroneous visual indicators outside the body, embedded markup symbols, imaging artifacts, etc.)
COVID-Net is by no means a production-ready solution. The team hopes, however, that researchers and citizen data scientists build upon the promising results achieved by the open-source COVID-Net on the COVIDx test dataset to accelerate the development of highly accurate, practical deep learning solutions for detecting COVID-19 cases from chest radiography images and accelerate treatment of those who need it most. Models and datasets will change over time as more research is conducted, and more COVID-19 X-ray images become available.