Previous methods using area scan and line scan cameras in DNA sequencing proved insufficient. Time delay and integration (TDI) cameras, however, amplify camera sensitivity to detect weak signals without degrading image quality by oversampling a scene across more rows of pixels, synchronized in a time-delayed fashion with the relative motion between the scene and line sensors. Adding the charges from multiple line sensors increases the signal-to-noise ratio, producing images with sufficient contrast at high speeds.
DNA sequencing, the process of determining the physical order of the four bases of DNA - A, G, C, and T, is a method for obtaining biometric data. Analyzing nucleic acid sequences can allow scientists to explore why a plant or animal exhibits the characteristics it exhibits and where it originally came from on Earth. The dramatically decreased cost of modern DNA sequencing, compared to the price for this testing in the early 2000s, has led to the widespread use of this testing in many scientific disciplines.
In order to conduct these tests, DNA is purified, cut into small pieces, and then loaded onto a special chip for analysis. Each of the four nucleotides is marked with a different fluorescent dye, which allows the researcher to differentiate the four nucleotides, A, G, C, and T. However, these dyes can appear very dim and need to be amplified using the polymerase chain reaction (PCR) method (bit.ly/VSD-PCR).
Related: Convolutional neural network detects COVID-19 from chest radiography images
Area cameras were used first for DNA sequencing. The camera captured dim fluorescent signals made on chips for the amplification and sequencing processes. Four different bases labeled with different fluorescence dyes enter the chip one by one in each round. The polymerase assembles a new DNA strand from fluorescently-labeled nucleotides in an order complementary to the purified DNA. Whenever the fluorescently-labeled nucleotide is individually inserted into the new DNA stand, the camera detects the fluorescence signal of the nucleotide and software determines the type of newly-inserted nucleotide by the wavelength of the fluorescent dye. Comparing and analyzing the base sequence of each piece of the purified DNAs sequentially connects the pieces of DNA, obtaining a complete DNA sequence.
Differentiating each nucleotide’s fluorescent signal from the image requires a high-magnification microscope because the several hundred million DNA pieces are planted on one chip, which measures several square centimeters. Therefore, it takes hundreds to thousands of shots to capture the entire chip at high magnification. This method took a significant amount of time and effort. To address this, scientific line scan cameras replaced area cameras and captured images at higher speeds. Detecting the fluorescent dye using line scan cameras required an image sensor with pixel sizes into dozens of µm, which led to low resolution images. Enter cameras, which offer higher sensitivity and speeds, and make for an even better option for DNA sequencing.
TDI line scan cameras use the time delayed integration technique, which use multiple stages to capture multiple exposures and generate cleared images. In each stage, photogenerated signal charges transfer in sync with object motion. Vieworks TDI cameras, for example, feature a hybrid image sensor with CCD pixel structure implemented by using CMOS technology, enabling high-speed image captures with up to 256 times great sensitivity, as a result of the cameras having up to 256 stages.
Bidirectional scanning represents another method for performing DNA sequencing faster. The dim fluorescent signal size of fluorescent dyes used for DNA sequencing can only be amplified so much, however, so a camera for DNA sequencing requires a certain level of noise and sensitivity. Specifically, the camera must be able to capture approximately 30 photon signals per one pixel and the light level difference between the background and the brightest pixel can be different by a factor of 16. When taking an image of an object with a TDI camera, its sensitivity can vary slightly depending on its moving direction. This may not be a problem in general applications, but slight differences may have larger consequences in DNA sequencing. Vieworks’ bidirectional dark signal non-uniformity (DSNU) ensures stability in imaging performance, regardless of direction (Figure 1).
As cameras used for DNA sequencing need to capture dim signals at high gain, bidirectional dark signal non-uniformity (DSNU) correction is needed. A lack of real-time DSNU correction requires manually conducting the correction process each time the direction is changed, which reduces the scanning speed. Vieworks’ real-time DSNU solves this problem by ensuring stability in imaging performance regardless of direction and ensuring fast scanning speed. As a result, Viework’s image sensors can capture images of the fluorescent substance with performance and dark correction technologies.
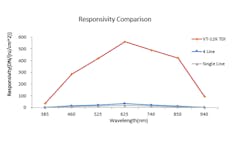
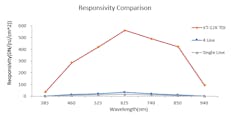
Internal company tests show the performance of a Vieworks TDI camera with CoaXPress interface compared to line scan cameras with one or four lines (Figure 2). At 385 and 940 nm wavelengths the performance of the TDI camera was slightly better, while the TDI camera’s performance was clearly superior for all wavelengths in-between. Responsivity is defined as the output in digital numbers (DN), aided by an input—areal energy density in nanojoules (nJ) / cm2.
About the Author

James Carroll
Former VSD Editor James Carroll joined the team 2013. Carroll covered machine vision and imaging from numerous angles, including application stories, industry news, market updates, and new products. In addition to writing and editing articles, Carroll managed the Innovators Awards program and webcasts.